Cellulose, which helps give plant cell walls their rigid structure, holds promise as a renewable raw material for biofuels — if researchers can accelerate the production process. Compared to the breakdown of other biofuel materials like corn, breaking down cellulose is slow and inefficient but could avoid concerns around using a food source while taking advantage of abundant plant materials that might otherwise go to waste. New research led by Penn State investigators has revealed how several molecular roadblocks slow this process.
The team’s most recent study, published in the Proceedings of the National Academy of Sciences, describes the molecular process by which cellobiose — a two-sugar fragment of cellulose that is made during cellulose deconstruction — can clog up the pipeline and interfere with subsequent cellulose breakdown.

Biofuel production relies on the breakdown of compounds like starch or cellulose into glucose, which can then be efficiently fermented into ethanol for use as a fuel or converted into other useful materials. The predominant biofuel option on the market today is generated from corn, in part because, the researchers said, their starches break down easily.
“There are several concerns about using corn as a biofuel source, including competing with the global food supply and the large quantity of greenhouse gasses produced when generating corn-based ethanol,” said Charles Anderson, professor of biology in the Penn State Eberly College of Science and an author of the paper. “A promising alternative is to break down cellulose from the non-edible parts of plants like corn stalks, other plant waste like forestry residue, and potentially dedicated crops that could be grown on marginal land. But one of the major things holding back so-called second-generation biofuels from being economically competitive is that the current process to break down cellulose is slow and inefficient.”
We have been using a relatively new imaging technique to explore the molecular mechanisms that slow down this process.”
Cellulose is composed of chains of glucose, held together by hydrogen bonds into crystalline structures. Scientists use enzymes called cellulases, derived from fungi or bacteria, to break down plant material and extract the glucose from the cellulose. But, the researchers said, cellulose’s crystalline structure paired with other compounds called xylan and lignin — also present in cell walls — provide additional challenges to the cellulose breakdown. Traditional techniques, however, were unable to reveal the specific molecular mechanisms of these slowdowns.
To explore these unclear mechanisms, the researchers chemically tagged individual cellulases with fluorescent markers. They then used Penn State’s SCATTIRSTORM microscope, which the team designed and built for this very purpose, to trace the molecules through each step of the breakdown process and interpreted the resulting videos using computational processing and biochemical modeling.
“Traditional methods observe the breakdown process at a larger scale, artificially manipulate the position of the enzyme or only capture molecules in motion, which means you may miss some of the naturally occurring process,” said Will Hancock, professor of biomedical engineering in the Penn State College of Engineering and an author of the paper. “Using the SCATTIRSTORM microscope, we were able to watch individual cellulase enzymes in action to really get at what is slowing down this process and generate new ideas for how to make it more efficient.”
The researchers specifically studied the effect of a fungal cellulase enzyme called Cel7A. As part of the breakdown process, Cel7A feeds cellulose into a sort of molecular tunnel, where it is chopped up.
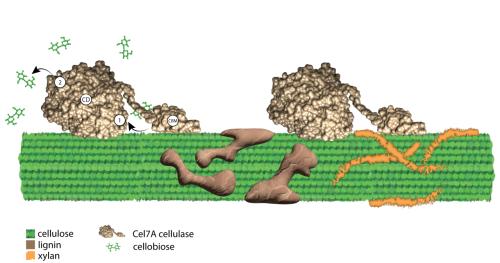
“Cel7A moves the glucose chain to the ‘front door’ of the tunnel, the chain is cleaved, and the products come out the ‘back door’ in a sort of pipeline,” said Daguan Nong, assistant research professor of biomedical engineering in the Penn State College of Engineering and first author of the paper. “We aren’t exactly sure how the enzyme threads the glucose chain to the tunnel or what exactly goes on inside, but we knew from previous studies that the product that comes out the back door, cellobiose, can interfere with the processing of subsequent cellulose molecules. Now, we know more about how it is interfering.”
Within the tunnel, Cel7A chops up cellulose — which has repeating units of glucose — into two-sugar cellobiose fragments. The researchers found that cellobiose in solution can bind to the “back door” of the tunnel, which can slow down the exit of subsequent cellobiose molecules as it essentially blocks the way. Additionally, they found that it can bind to Cel7A near the front door, preventing the enzyme from binding to additional cellulose.
“Because cellobiose is so similar to cellulose, it’s maybe not surprising that the little pieces can get into the tunnel,” Hancock said. “Now that we have a better understanding of how exactly cellobiose is mucking things up, we can explore new ways to fine tune this process. For example, we could alter the front or the back door of the tunnel or change aspects of the Cel7A enzyme to be more efficient at preventing this inhibition. There has been a lot of work to engineer more efficient cellulase enzymes over the last two decades, and it’s an incredibly powerful approach. Having a better understanding of the molecular mechanisms that limit cellulose degradation will help us direct this effort.”
This research builds off recent work by the research team to understand other roadblocks to the degradation process — xylan and lignin — which they published recently in RSC Sustainability and Biotechnology for Biofuels and Bioproducts.
“We found that xylan and lignin operate in different ways to interfere with the breakdown of cellulose,” said Nerya Zexer, postdoctoral researcher in biology in the Penn State Eberly College of Science and lead author of the RSC Sustainability paper. “Xylan coats the cellulose, reducing the proportion of the enzymes that can bind to and move cellulose. Lignin inhibits the enzyme’s ability to bind to cellulose as well as its movement, reducing the velocity and distance of the enzyme.”
Although strategies exist to remove components like xylan and lignin from the cellulose, the researchers said the removal of cellobiose is more difficult. One method uses a second enzyme to cleave cellobiose, but it adds additional cost and complexity to the system.
“About 50 cents per gallon of bioethanol production costs is dedicated just to enzymes, so minimizing this cost would do a lot in terms of making bioethanol from plant waste more competitive with fossil fuels or corn-based ethanol,” Anderson said. “We will continue to investigate how to engineer enzymes and explore how enzymes might work together with the goal of making this process as low-cost and efficient as possible.”
The research team at Penn State also includes Zachary Haviland, undergraduate student majoring in biomedical engineering at the time of the research; Sarah Pfaff, graduate student in biology at the time of the research; Daniel Cosgrove, Holder of the Eberly Family Chair in Biology; Ming Tien, professor emeritus of biochemistry and molecular biology; and Alec Paradiso, undergraduate student majoring in biotechnology.
Funding from the U.S. Department of Energy (DOE) and the U.S. National Science Foundation supported this work, including the construction of the SCATTIRSTORM microscope. Additional support was provided by the Center for Lignocellulose Structure and Formation, an Energy Frontier Research Center funded by the DOE.
